Grounded Visual Intelligence
The MedGemma MRI Pipeline is meticulously engineered to support anatomically grounded MRI interpretation for highly complex glioblastoma analysis. By seamlessly integrating localized automated tumor segmentation logic built natively over standardized anatomical coordinates, the system achieves perfect resonance with atlas space geometry.
Unlike purely generative black-box frameworks, MedGemma explicitly extracts structured anatomical facts before passing verified findings into an interactive language-enabled reasoning layer. This structural grounding ensures maximum hallucination reduction and peerless transparency when users execute the pipeline.
Processing Pipeline Workflow
Simply provide multimodal MRI structural scans (T1, T1CE, T2, FLAIR). The pipeline rigidly executes four critical deterministic stages locally on the hardware before exposing the high-performance visualization dashboard.
Segmentation
Precisely isolate the glioblastoma boundaries and distinct tumor subregions directly from the raw NIfTI datasets.
Atlas Registration
Perform rigid mapping of the scan into physical MNI atlas space, ensuring the morphology aligns against normalized geometry.
Fact Extraction
Derive structural conclusions. Extract factual laterality metrics, lobar involvement, and precise volumetric intersection capacities natively.
Language Grounding
Deploy a localized large vision-language model architecture to generate clinical summaries directly anchored to structural evidence.
System Flow Diagram
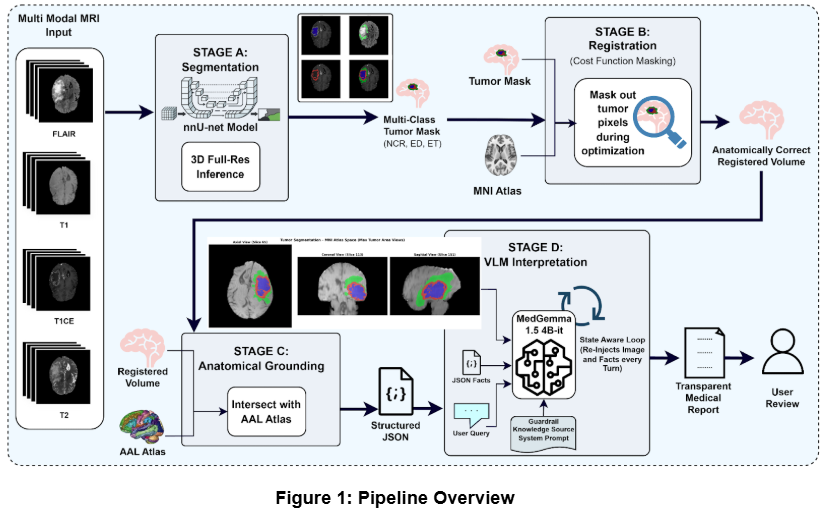
Figure 1. Architectural blueprint of the deterministic MedGemma multimodal pipeline infrastructure.
Interactive Clinical UI
The system features a bespoke, offline-first dashboard designed to seamlessly integrate deterministic data processing with natural language interaction.
Subject Library & Deterministic Anatomical Grounding
Navigate a localized clinical database of evaluated subjects, ranked immediately by AI confidence scores. Selecting a patient instantly surfaces their metadata, extracted deterministically rather than probabilistically. Following a rigorous Anatomical Atlas (AAL) grounding phase, the pipeline intersects the registered tumor mask with the atlas to calculate precise volumetric capacities across anatomical structures, such as the Temporal or Occipital Lobes. These deterministic features, including laterality and midline crossing, are compiled into a structured "Truth Anchor," serving as the immutable factual basis for all downstream analysis.
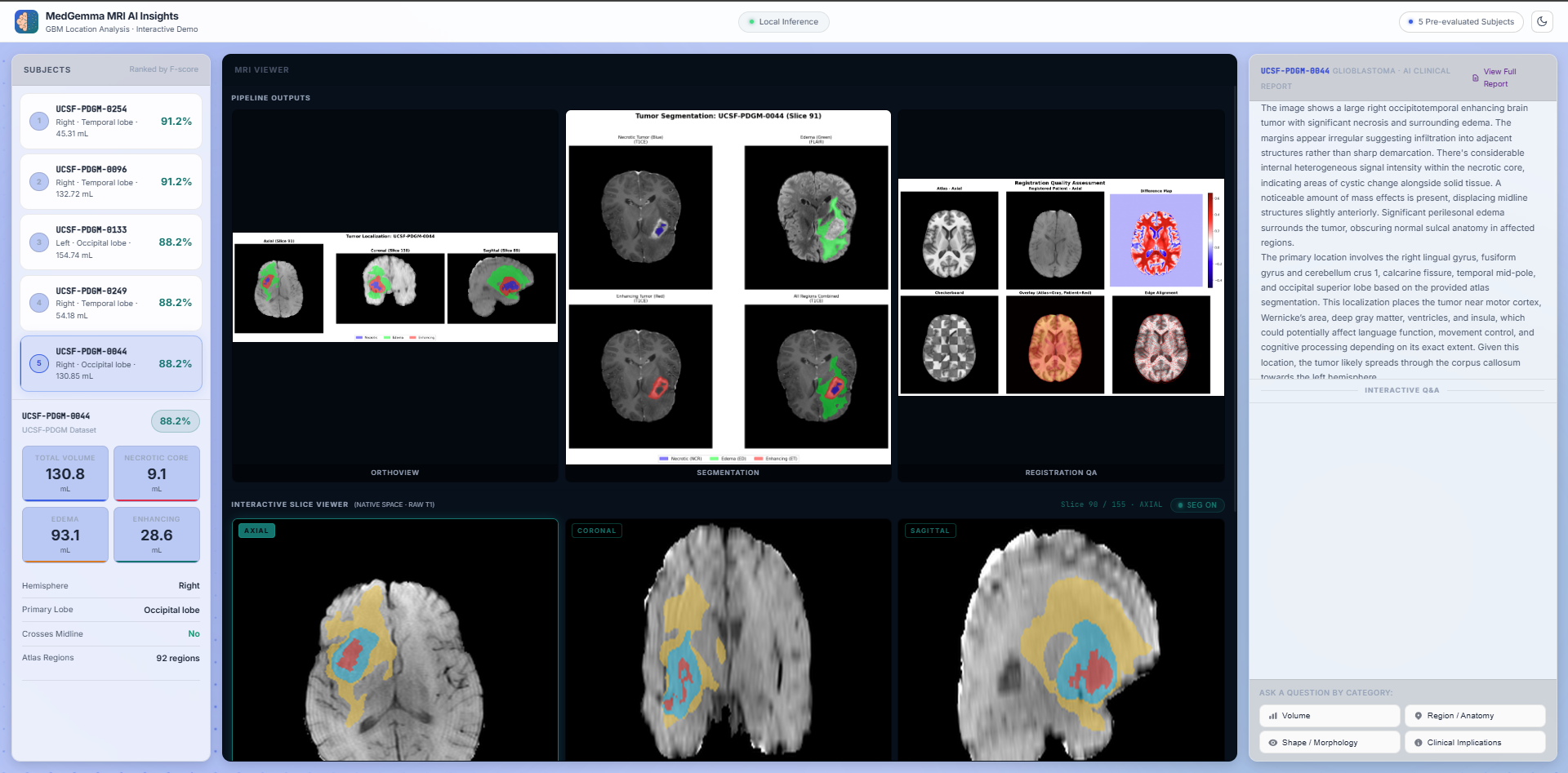
Native Visual Grounding & Agentic Chat
Explore high-contrast NIfTI MRI slices dynamically while interacting with the constrained MedGemma 1.5 reasoning engine. The AI seamlessly interprets complex morphological characteristics (like necrotic cores and edema margins) by directly referencing deterministic pipeline outputs. To eliminate dangerous clinical hallucinations and multi-turn context drift, the interface employs a continuous state-aware loop known as the Anti-Amnesia protocol. By dynamically re-injecting the Truth Anchor into the model's context window at every conversational turn, the system maintains strict spatial fidelity throughout prolonged clinical inquiries.
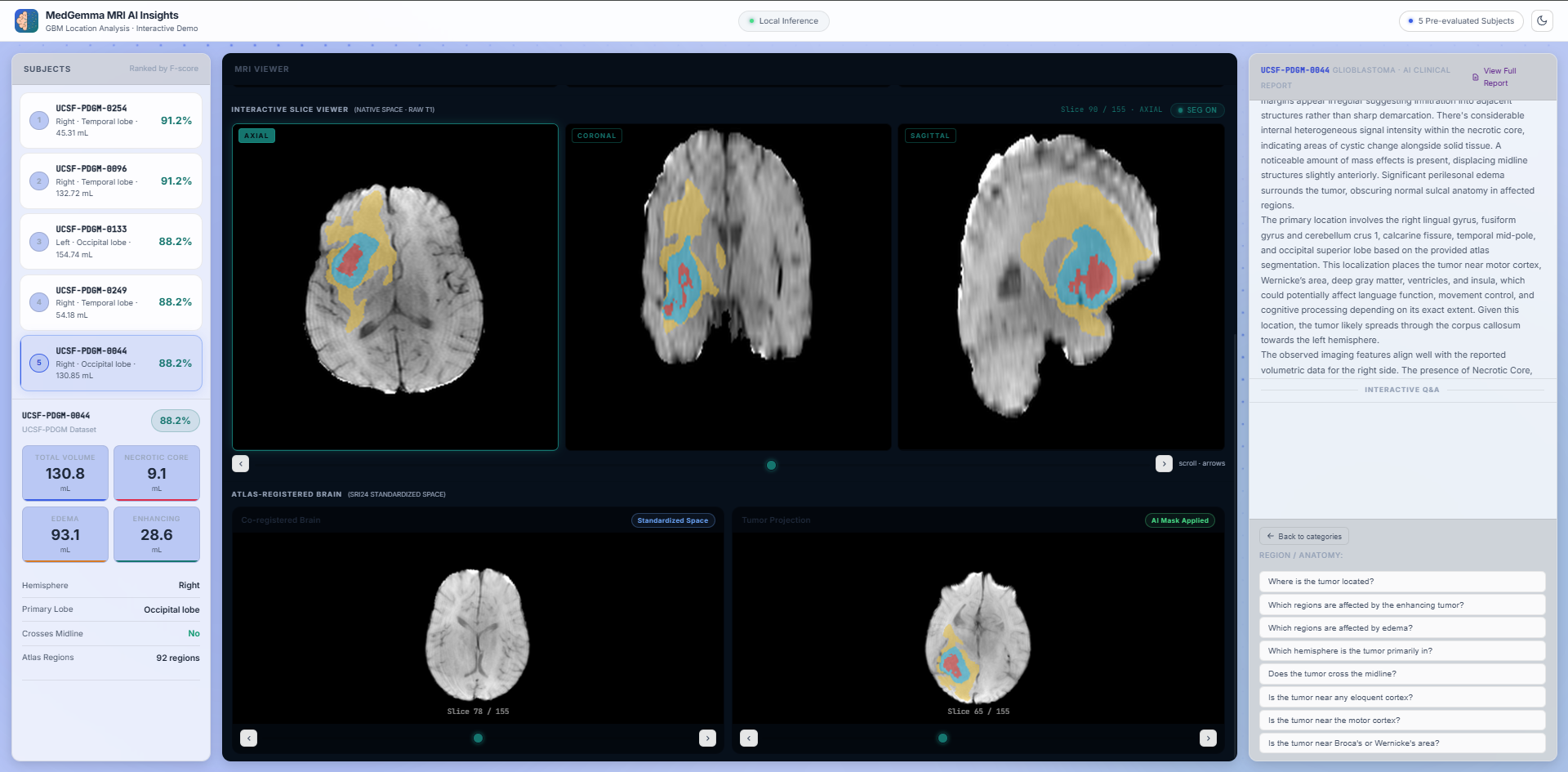
Pipeline Outputs: Structure-Preserving Normalization
View the raw, intermediate, and final pipeline outputs with complete transparency. The visualizer highlights the initial segmentation achieved via a 3D U-Net architecture (nnU-Net v2) operating on multimodal MRI sequences. Crucially, the system displays the Registration Quality Assessment, demonstrating the application of Cost Function Masking (CFM) during symmetric diffeomorphic registration (ANTs SyN). By actively ignoring tumor pixels during optimization, CFM prevents the lesion from artificially distorting to fit the healthy reference template, ensuring highly accurate topological preservation.